This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison. https://genetics564.weebly.com/
What Are Post-Translational Modifications?
|
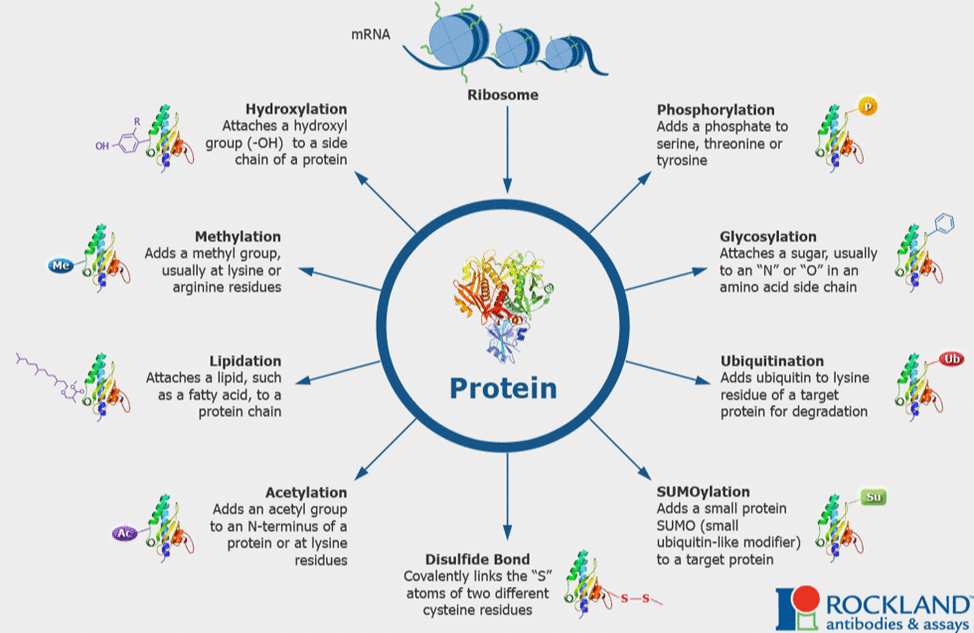
Post-translational modifications are any changes to a protein after they are translated. These can have wide effects on the cell and can happen at anytime post translation. Additionally, these modifications can vary from cell type to cell type. These can be markers for signaling or changes needed for pathways (phosphorylation), peptide breakdown (proteolysis), markers for degradation (ubiquitination), methylation, or addition of fats or sugars (lipidation and glycosylation). All types of modifications are pictured on the left. These can change at different cell stages, so analysis at different stages are crucial. Overall, these changes are important to understand because they can have huge effects on the function of a protein and their respective protein networks and interactions. [1,2] |
How Are These Modifications Detected?
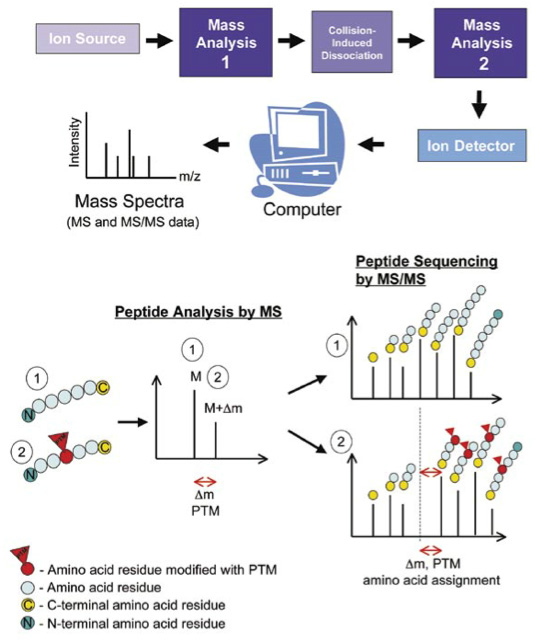
Proteins can be marked at different stages of protein analysis (Figure 2). Then, their relative weight will be different because they have an addition or have been cleaved. This can be seen through mass spectrometry.
How Were Post-Translational Modifications Detected in DBT?
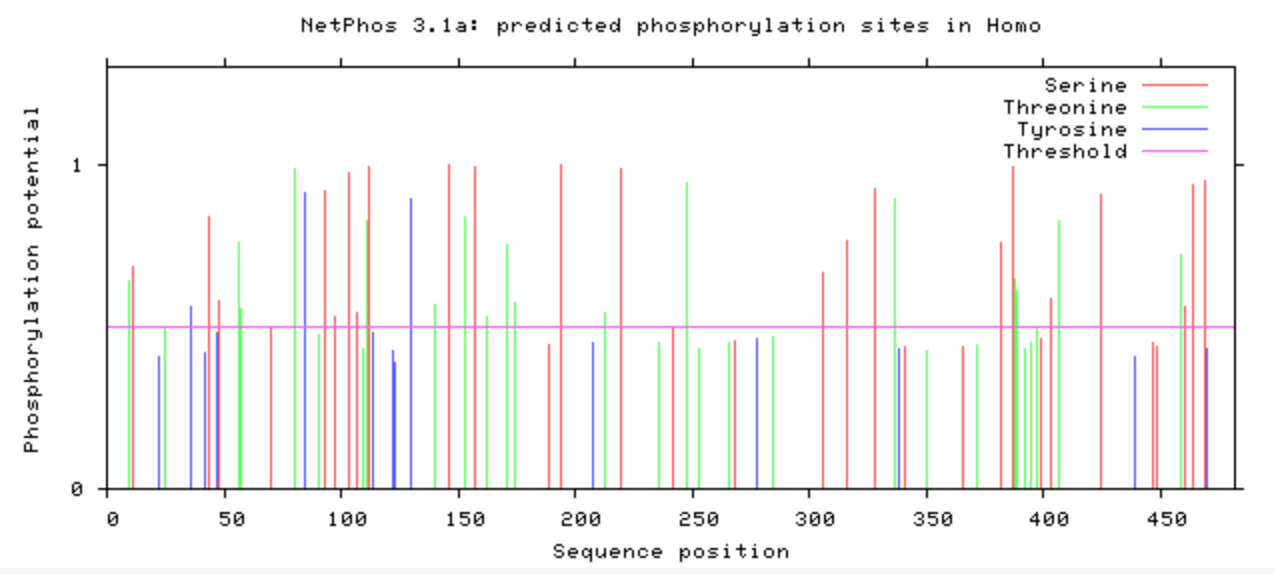
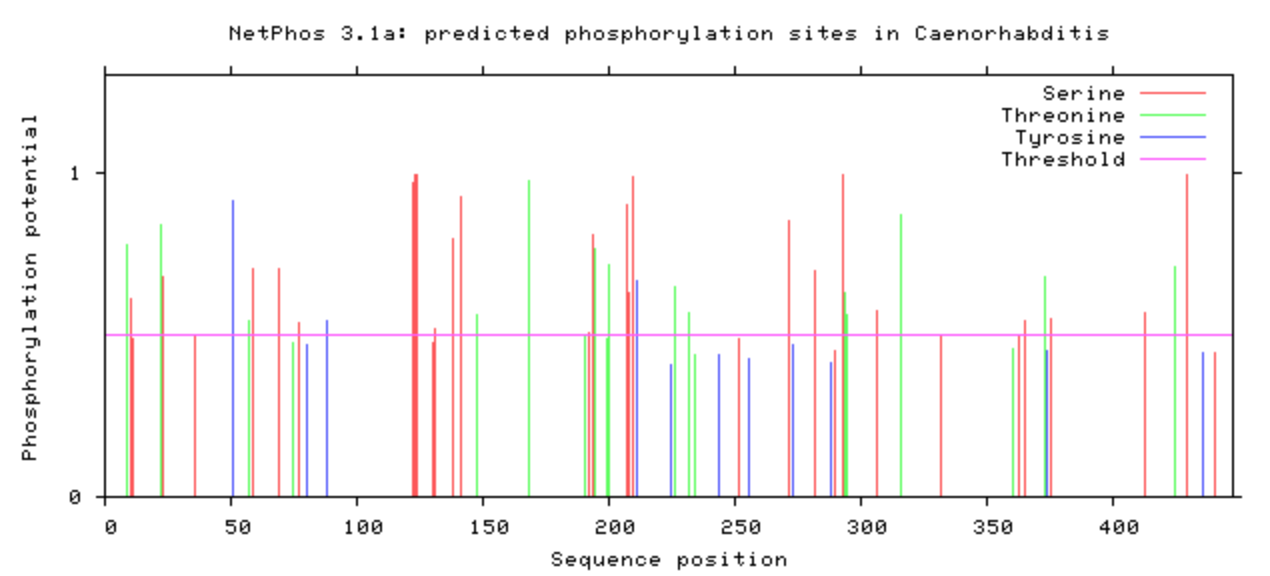
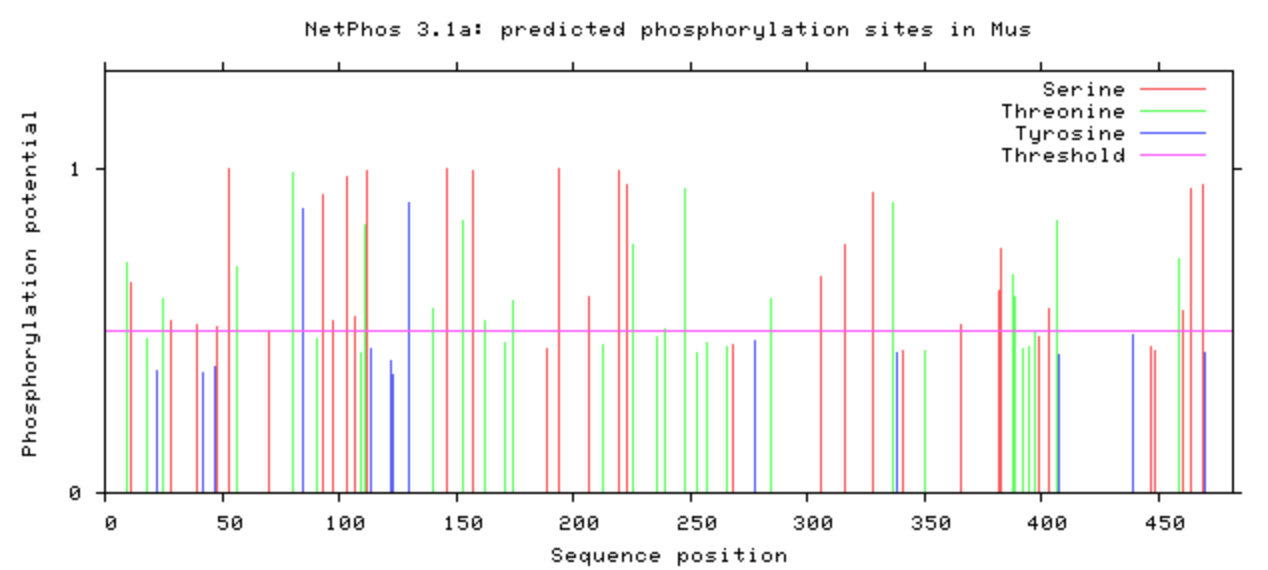
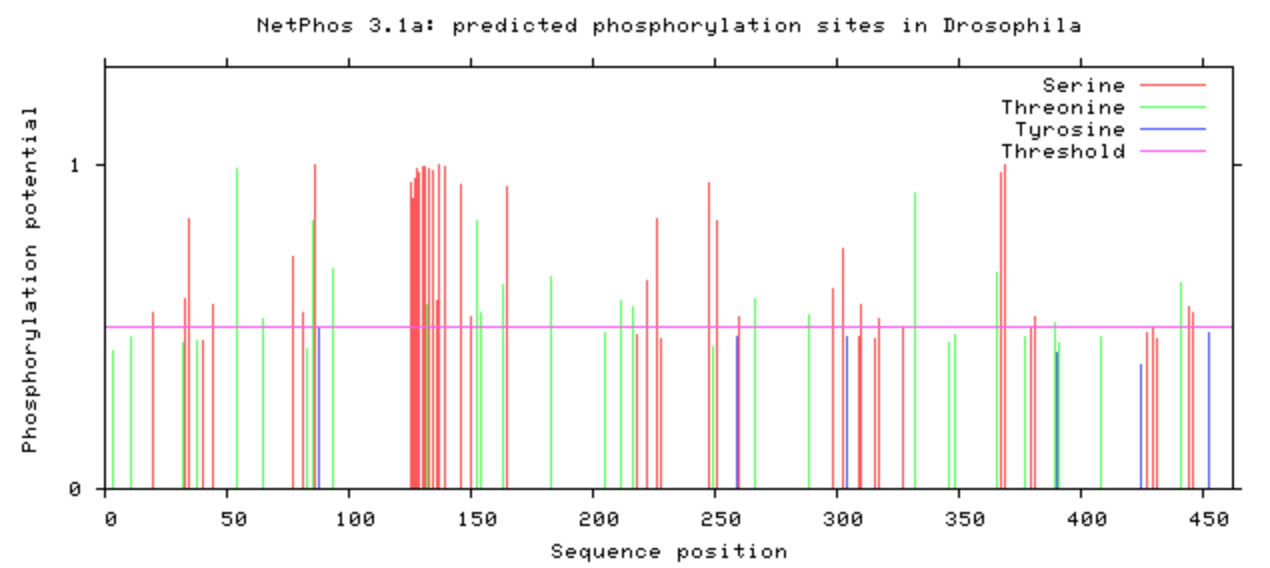
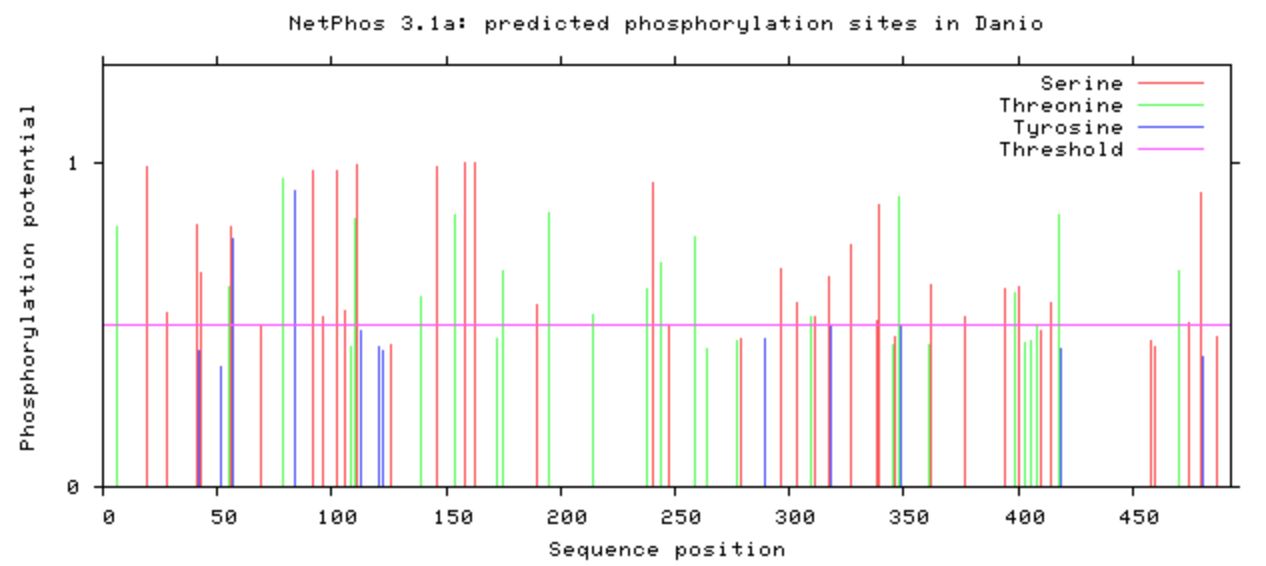
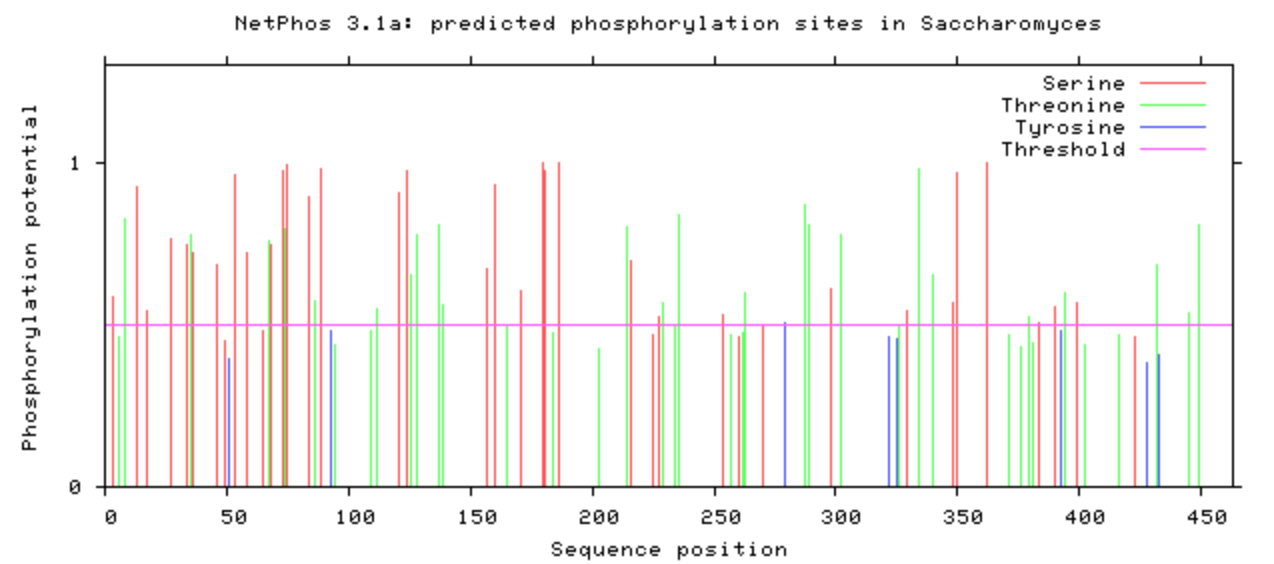
NetPhos was used to computational determine what the post-translational modifications were on the DBT protein.
Discussion
The phosphorylation was looked at in all model organisms, particularly the human and mouse. This information can be used when determining what is conserved among species and indicate what may be affected upon mutation with CRISPR/Cas9. If it is phosphorylated, then mutating it to be an amino acid that will not be phosphorylated may have additional affects that may be of interest. A particular amino acid of interest was the human 213 amino acid, which is conserved among all complex digestive system organisms. Additionally, it is phosphorylated among all complex organisms (except zebrafish). This could indicate variants in this amino acid may contribute to the poor feeding phenotype.
References
Sources:
[1] Overview of Post-Translational Modifications (n.d) Retrieved from: https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
[2] Pascovici, D., Wu, J., et al. (2019). Clinically Relevant Post-Translational Modification Analyses - Maturing Workflows and Bioinformatics Tools. Retrieved from: https://www.mdpi.com/1422-0067/20/1/16/htm
Images:
Header: 182-InsulinReceptor_1irk_1ir3_JSmol.jpg
PTM of Homologs: www.cbs.dtu.dk/services/NetPhos/
All other images linked
[1] Overview of Post-Translational Modifications (n.d) Retrieved from: https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
[2] Pascovici, D., Wu, J., et al. (2019). Clinically Relevant Post-Translational Modification Analyses - Maturing Workflows and Bioinformatics Tools. Retrieved from: https://www.mdpi.com/1422-0067/20/1/16/htm
Images:
Header: 182-InsulinReceptor_1irk_1ir3_JSmol.jpg
PTM of Homologs: www.cbs.dtu.dk/services/NetPhos/
All other images linked