This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison. https://genetics564.weebly.com/
What Are Protein Domains?
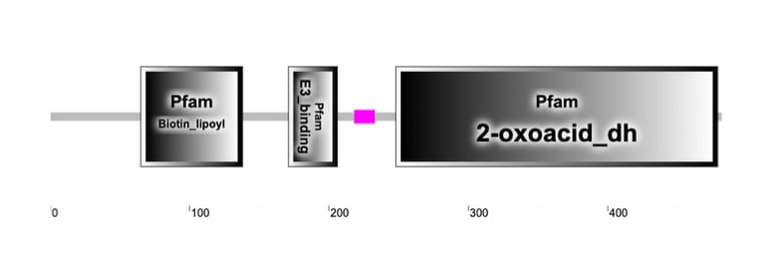
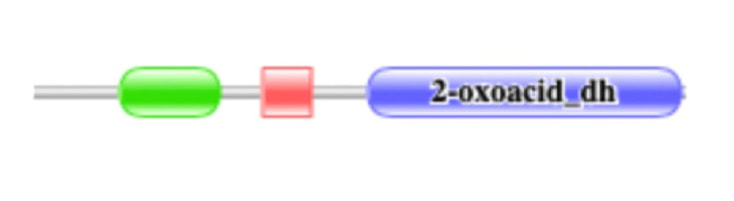
Protein domains are classified sequences on a protein hat have a specific structure or function. [1] These domains are frequently seen in other proteins and/or species that have a similar function. [2] Each domain can have a structural, catalytic, binding, or interactive function that are all necessary for the protein or protein complex to work correctly. The domains can be repeated within the protein and could be found in other proteins. This can be used to determine protein function, compare homologous proteins, or create protein families. [1] PFAM and SMART are two data bases that are used to determine the protein domains for humans and homologs of interest. Both were used to analyze the protein domains of the DBT protein (Figures 1A and B).
DBT Protein Domains
Protein Domain Analysis Among Homologs
Discussion
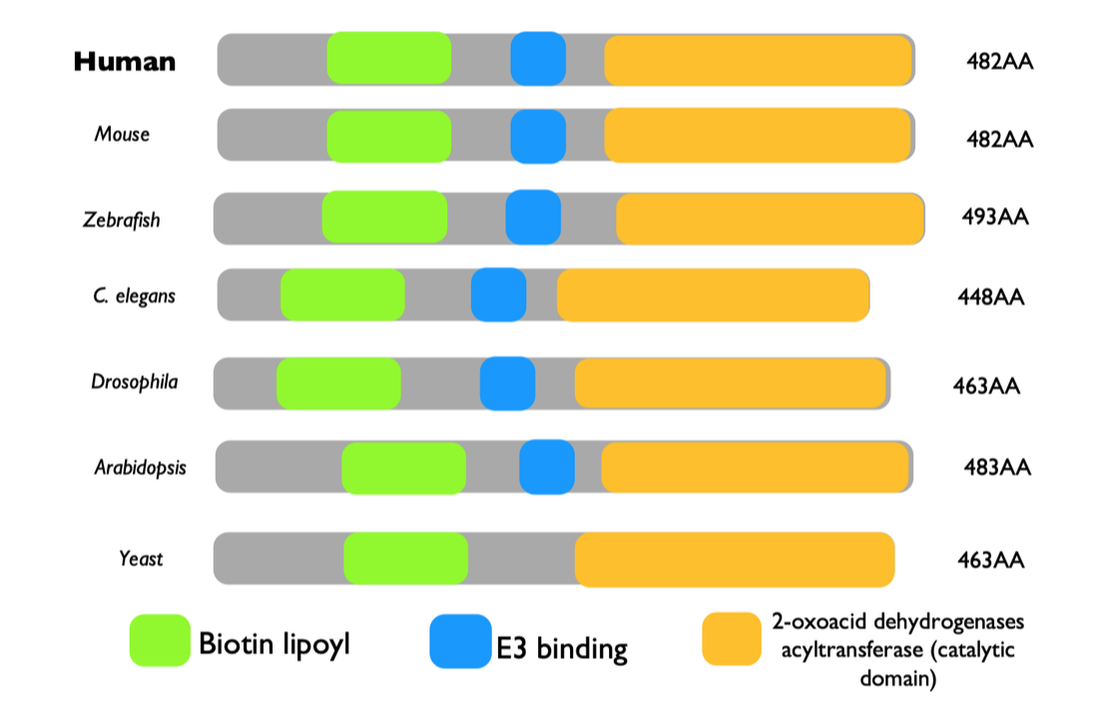
DBT was found to have three well characterized protein domains in both SMART and Pfam. These domains included the Biotin lipoyl domain, the E3 binding domain, and the 2-oxoacid dehydrogenases acytransferase domain (catalytic). Upon investigation through SMART and Pfam, common model organisms’ domains were also mapped on the amino acid sequence (shown above). All, except for yeast, were found to have the same domains of almost identical sequence length. Yeast was missing the E3 binding domain. This provides insight into what domains are highly conserved and what regions are necessary for function of this protein. Gene Ontology confirms that there is conservation of the function, which would also likely require conservation of essential domains. Many of the pathogenic or likely pathogenic variants are in these domains, but there are regions outside of the domains that contain pathogenic variants. [5] This could be an area of further research.
References:
[1] EMBL-EBI. (n.d.) Family- and Domain-based protein classification. Retrieved from: https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/family-and-domain-based-protei
[2] ProteinStructures.com. (n.d.) Protein Domains and Domain Classification. Retrieved from: https://proteinstructures.com/Structure/Structure/protein-domains.html
[3] EMBL. (2019). Domains within Homo sapiens protein ODB2_HUMAN (P11182). Retrieved from: http://smart.embl-heidelberg.de/smart/job_status.pl?jobid=72332246159981550784640ApouZdsfcO#
[4] EMBL-EBI. (n.d.) 2 oxoacid dehydrogenase acyltransferase, catalytic domain. Retrieved from: https://www.ebi.ac.uk/interpro/entry/IPR001078
[5] ClinVar. NCBI. Retrieved from https://www.ncbi.nlm.nih.gov/clinvar/?term=dbt%5Bgene%5D
Images:
Header: https://pm1.narvii.com/6980/fb8c96c73493899db5f67dd62aeff641b1420b88r1-800-612v2_hq.jpg
All other figures are linked
[1] EMBL-EBI. (n.d.) Family- and Domain-based protein classification. Retrieved from: https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/family-and-domain-based-protei
[2] ProteinStructures.com. (n.d.) Protein Domains and Domain Classification. Retrieved from: https://proteinstructures.com/Structure/Structure/protein-domains.html
[3] EMBL. (2019). Domains within Homo sapiens protein ODB2_HUMAN (P11182). Retrieved from: http://smart.embl-heidelberg.de/smart/job_status.pl?jobid=72332246159981550784640ApouZdsfcO#
[4] EMBL-EBI. (n.d.) 2 oxoacid dehydrogenase acyltransferase, catalytic domain. Retrieved from: https://www.ebi.ac.uk/interpro/entry/IPR001078
[5] ClinVar. NCBI. Retrieved from https://www.ncbi.nlm.nih.gov/clinvar/?term=dbt%5Bgene%5D
Images:
Header: https://pm1.narvii.com/6980/fb8c96c73493899db5f67dd62aeff641b1420b88r1-800-612v2_hq.jpg
All other figures are linked