This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison. https://genetics564.weebly.com/
What Are Protein Interaction Networks?
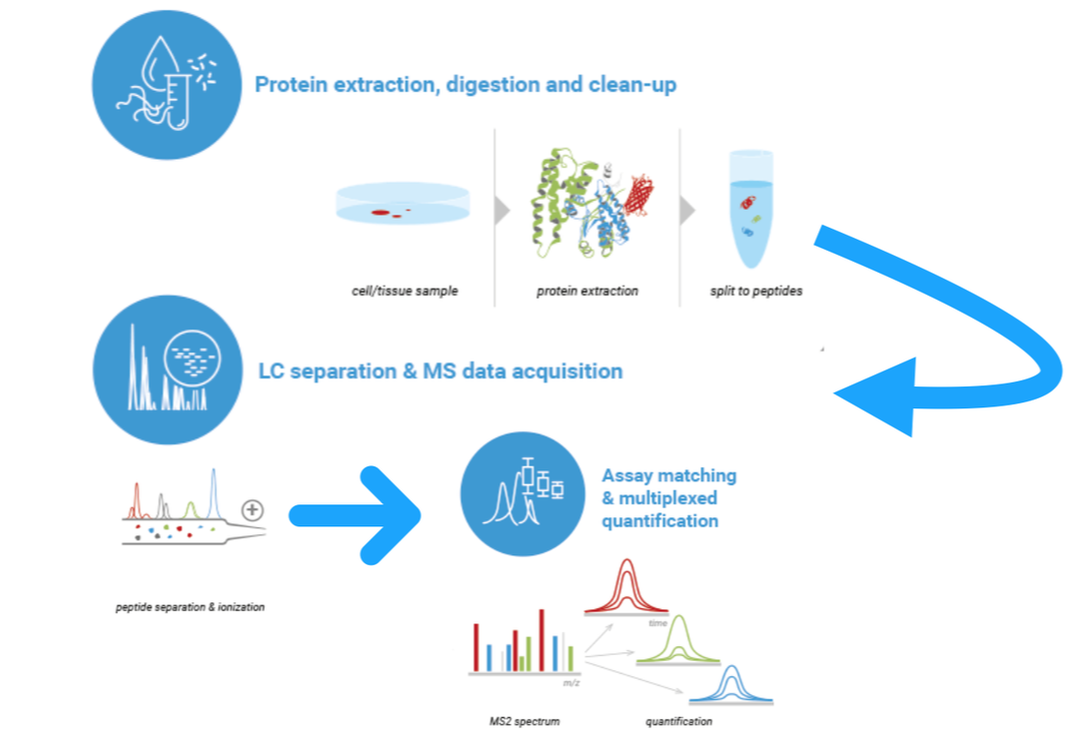
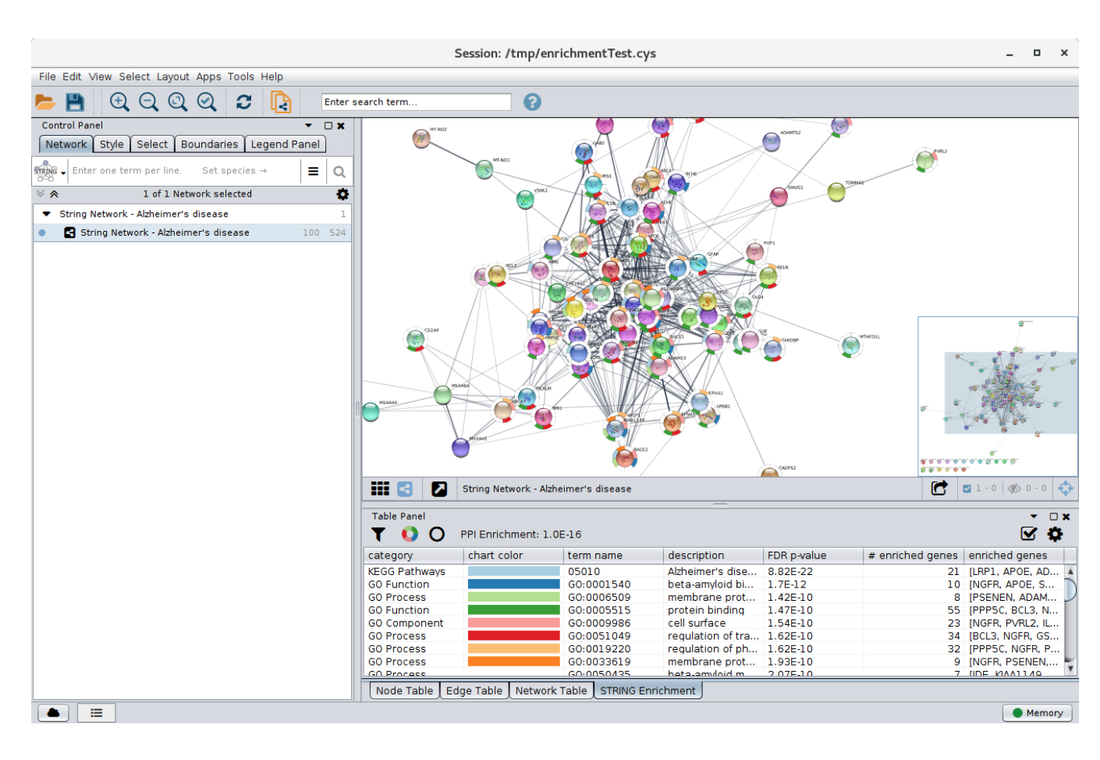
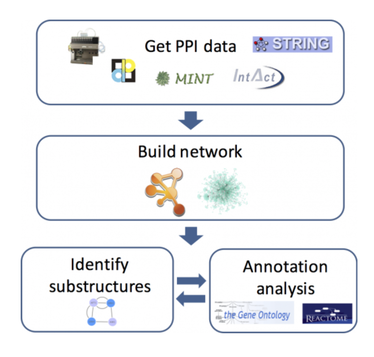
Protein interaction networks depict how proteins interact with other proteins in cells or body systems. They can indicate that those proteins are part of a signaling cascade, bind to each other, or affect the overall function of each other. The interactions can either be dynamic or constant. These protein interaction networks can be determined experimentally (Figure 1A) or through computer programs such as STRING (Figure 1B). Computer programs show experimentally determined or predicted interactions. All of this information can be used to develop a complex protein interaction network (Figure 2) that can then be sorted by pathway, function, or system. These interaction networks can be made for a specific protein of interest or entire cell protein interaction.
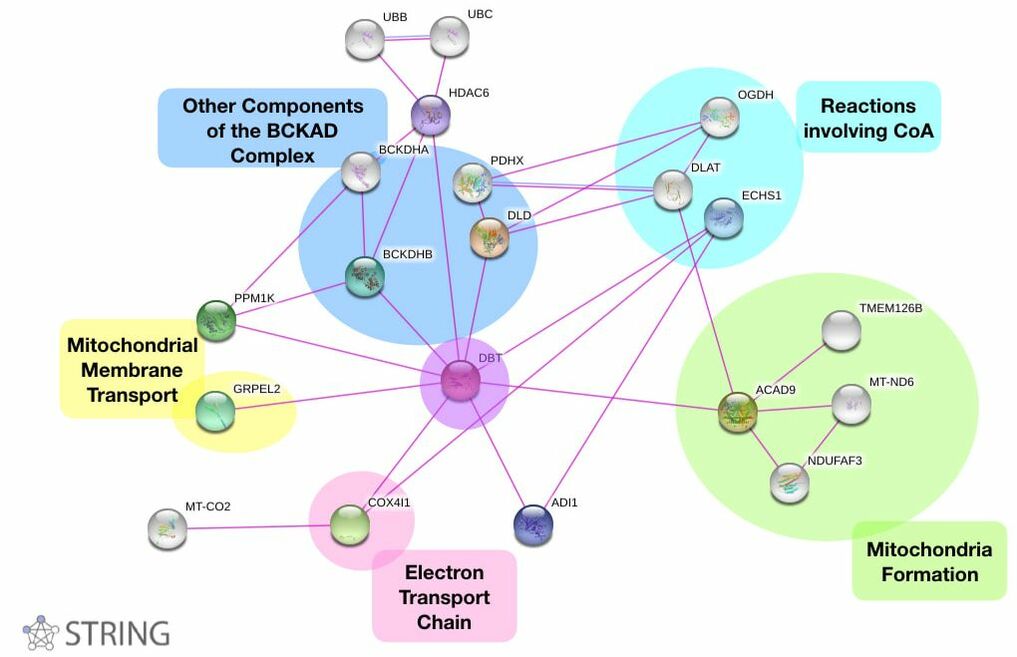
DBT Protein Interaction Network
Discussion
It was expected that DBT interacted with the other proteins in the branched chain alpha-ketoacid dehydrogenase complex. However, there were other proven direct interactions with other proteins of the mitochondria. All of the gene ontologies are indicated in the map above. The protein interaction with GREPL2 was particularly of interest because it highly expressed in the esophagus. [3] This was intriguing because Maple Syrup Urine Disease is known to have a poor phenotype. Other mitochondrial disorders have been associated with GERD in infants that results in poor feeding [4,5]. This association and gap in knowledge was used in this study's specific aims. Additionally, there is information on what other functions are impacted by DBT. Mutations may disrupt all of these functions in various ways. These are all areas that could be further researched in the future.
References:
Sources:
[1] Protein-protein interaction networks. (n.d.) EMBL-EBI. Retrieved from: https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
[2] Snider, J., Kotlyar, Max., et al. (2015). Fundamental of Protein Interaction Network Mapping. Retrieved from: http://msb.embopress.org/content/11/12/848#abstract-2
[3] https://www.ncbi.nlm.nih.gov/gene/134266
[4] Finsterer, J., & Frank, M. (2016). Gastrointestinal manifestations of mitochondrial disorders: a systematic review. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5330602/
[5] Gouspillou Gilles, Hepple Russell T. (2016). Mitochondria in Skeletal Muscle Health, Aging and Diseases. Retrieved from: https://www.frontiersin.org/articles/10.3389/fphys.2016.00446/full
Images:
Header: The_protein_interaction_network_of_Treponema_pallidum.png
All other images linked
[1] Protein-protein interaction networks. (n.d.) EMBL-EBI. Retrieved from: https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
[2] Snider, J., Kotlyar, Max., et al. (2015). Fundamental of Protein Interaction Network Mapping. Retrieved from: http://msb.embopress.org/content/11/12/848#abstract-2
[3] https://www.ncbi.nlm.nih.gov/gene/134266
[4] Finsterer, J., & Frank, M. (2016). Gastrointestinal manifestations of mitochondrial disorders: a systematic review. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5330602/
[5] Gouspillou Gilles, Hepple Russell T. (2016). Mitochondria in Skeletal Muscle Health, Aging and Diseases. Retrieved from: https://www.frontiersin.org/articles/10.3389/fphys.2016.00446/full
Images:
Header: The_protein_interaction_network_of_Treponema_pallidum.png
All other images linked